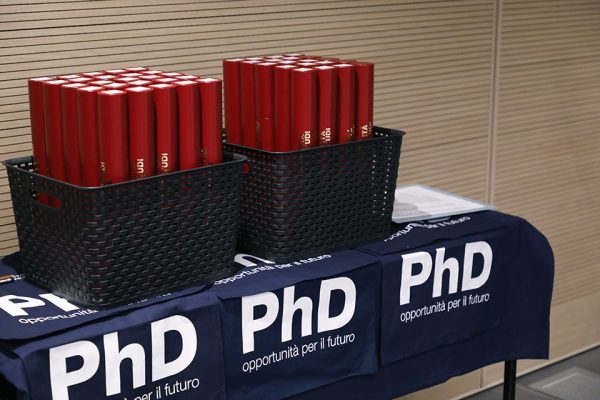
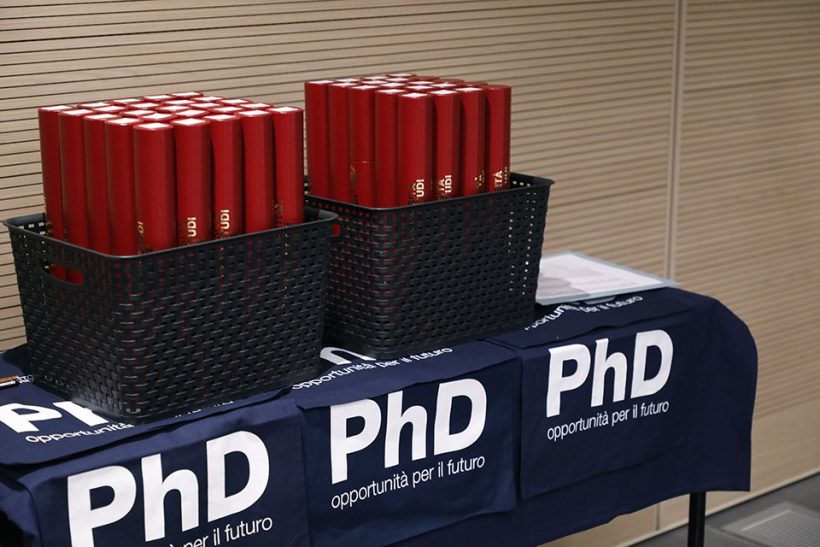
The Computational Biophysics Group at the University of Udine, Italy, invites applications for a number of PhD fellowships in computational biophysics applied to nanomedicine. Qualified candidates are encouraged to reach out to prospective supervisors in advance to discuss possible projects to be included in the application package, which must be submitted to the PhD School […]
Author: admin
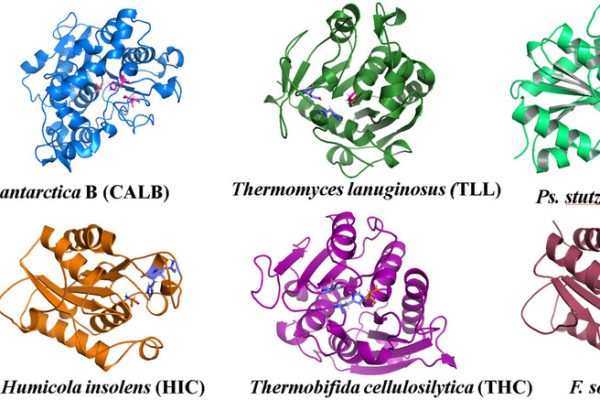
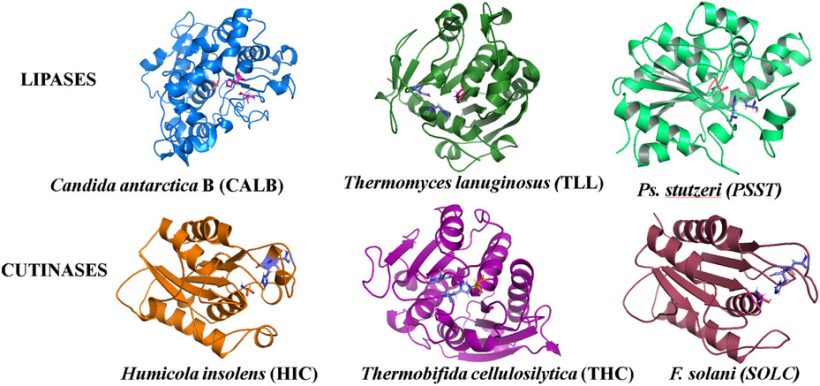
Enzymes for Synthesis and Degradation of Bio-Based Polyesters: Automatic In Silico Screening and Experimental Validation
Our study presents a computational workflow to support the eco-design of sustainable polymers by identifying effective biocatalysts for polyester synthesis and degradation. Using well established computational tools like Gromacs (molecular dynamics) and AutoDock (docking), integrated in a modeFRONTIER workflow, the pipeline evaluates enzyme–substrate interactions and ranks candidates based on predicted performance. Screening multiple bio-based monomers […]
Now at the Department of Mathematics, Computer Science and Physics at the University of Udine, Italy!
After a pause from Academia: I am back! I indeed just took up a position in Applied Physics at the Department of Mathematics, Computer Science and Physics at the University of Udine, Italy. My new coordinates: e-mail: sara.fortuna@uniud.it web: www.sarafortuna.eu Office: B1.2 (first floor) Dipartimento di Scienze Matematiche, Informatiche e Fisiche Universita’ di Udine Via […]
We are part of the H2020-MSCA-ITN team STOP SPREAD BAD BUGS!
We are part of the team being awarded the H2020 Marie Skłodowska-Curie Actions (MSCA) Innovative Training Networks (ITN) with the project “STOP SPREAD BAD BUGS: novel antimicrobial approaches to combat multidrug resistance in bacteria“. As part of the CONCEPT lab we will support a multidisciplinary team composed by medical doctors, biologists, chemists, and physicists to […]
We are part of the PRIN 2020 project CANNOT-ESKAPE!
We are part of the team being awarded the national PRIN2020 project “CANNOT-ESKAPE: Targeting baCteriAl cell eNvelope of Nocosomial paThogens to ESKAPE resistance”. Our unit, led by Rita de Zorzi (University of Trieste), we will support a multidisciplinary team led by Flavia Squeglia (CNR, Naples), supporting biologists, chemists, and physicists tackling the problem of antimicrobial […]
Read our latest paper on the topology to employ for in silico designed peptides!
A new application of BINDesignER/PARCE! Herein, we compared the ability of linear and cyclic peptides generated in silico to target different protein sites: internal pockets and solvent-exposed sites. We selected human lysozyme (HuL) as a model target protein combined with the computational evolution of linear and cyclic peptides. The computational results demonstrated that cyclic peptides […]
Welcome to Patricio Barletta, who joined today our AIRC research project.
We welcome Patricio Barletta in our team! Patricio got his PhD from the Universidad Nacional de Quilmes, Argentina in 2020 and joined our team in the framework of the ICTP-TRIL fellowship program. He will work on the computational aspects of our project. He will carry out his activity in the CONCEPT Lab head by Walter […]
We have been awarded a “ARDF’s Annual Open grant” from the Alternatives Research & Development Foundation (ARDF)
Our commitment in the development of new computational protocols that can compete and, in time, eliminate the demand for in vivo procedures for antibody discovery and maturation has been prized once again with a 40,000$ grant with the project ” De-novo engineered antibody fragments: validation of the first in-silico pipeline for antibody discovery ” from the […]
Welcome to Theo Battista, who joined today our AIRC research project.
We welcome Theo Battista in our team! Theo just got his PhD in Biochemistry from “Sapienza”, University of Rome. He will carry out his activity at the Structural Biology Lab head by Paola Storici at Elettra, Trieste, Italy.
Siamo stati in onda sul TG Leonardo!
Nell’ambito di una collaborazione con Trasactivia SrL e l’Universita’ Tor Vergata di Roma stiamo contribuendo allo sviluppo di un nuovo vaccino prodotto interamente dalle foglie delle piante di tabacco. Il progetto e’ stato presentato su Rai3 nell’ambito del TG Leonardo. Ci trovate al minuto 8.51 -11.31